New insights into how diet and medication impact the influence of gut bacteria on our health
Research published in Cell on 29th August by the groups of Filipe Cabreiro from the MRC London Institute of Medical Sciences and Imperial College and Christoph Kaleta from Kiel University in Germany has demonstrated that diet can alter the effectiveness of a type-2 diabetes drug via its action on gut bacteria.
Bacteria that reside in our gastrointestinal tract, referred to as the gut microbiome, produce numerous molecules capable of influencing health and disease. The function of the gut microbiome is known to be regulated by both diet and drugs such as the drug metformin, which is used to treat type-2 diabetes and has been shown to extend the lifespan of several organisms. However, understanding the complicated, multi-directional relationships between diet, drugs and the gut microbiome represents a considerable challenge. “Disentangling this network of interactions is of utmost importance since the specific mechanism of action of metformin is still unclear,” says Filipe Cabreiro.
You are currently viewing a placeholder content from YouTube. To access the actual content, click the button below. Please note that doing so will share data with third-party providers.
Video: © MRC London Institute of Medical Sciences
A new screening technique
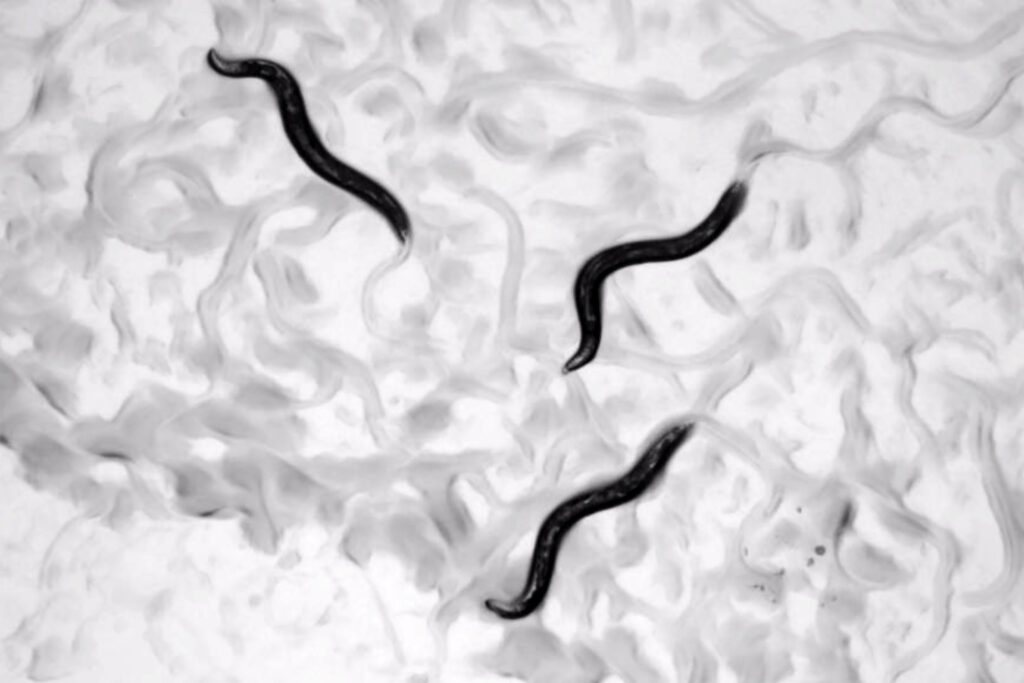
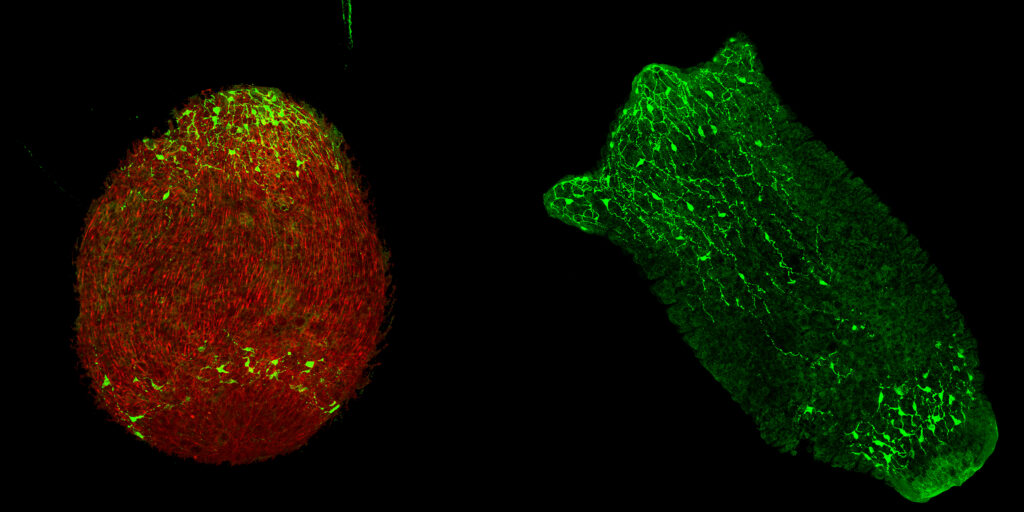
Cabreiro and his team developed an innovative four-way high-throughput screening technique to better understand how diet, drugs and the gut microbiome interact to influence host physiology. They used the nematode worm Caenorhabditis elegans colonized with the human gut bacteria Escherichia coli as a simplified host-microbiome model and exposed it to metformin in the presence of hundreds of different nutritional compounds.
They found that metformin treatment altered the metabolism and lifespan of the C. elegans host and that these effects could be either enhanced or suppressed by specific nutrients. Crucially, it was revealed that the gut bacteria played a key role in mediating this phenomenon.
The importance of the diet and gut bacteria explain why metformin was previously shown to have no effect on the lifespan of another commonly studied organism, the fruit fly. Helena Cochemé, who collaborated on this study says “As it turned out, the typical laboratory food of fruit flies is rich in sugars. After taking away the sugar we also saw positive effects of metformin in fruit flies colonized with E. coli.”
Bacterial nutrient signaling is a central modulator of microbe-host-drug interactions
Further analysis revealed that bacteria possess a sophisticated mechanism that enables them to coordinate nutritional and metformin signals and to rewire their own metabolism accordingly. As a result of this adaptation, the bacteria accumulate a metabolite called agmatine which was shown to be required for the positive effects of metformin on host health.
What about humans?
Cabreiro collaborated with Christoph Kaleta from Kiel University to investigate whether the results found in C. elegans could also be observed in the more complex microbiota of humans. They analysed data related to the microbiome, nutrition and medication status of a large cohort of type-2 diabetic patients and healthy controls. “Intriguingly, we found that metformin treatment was strongly associated with an increased capacity for bacterial agmatine production,” says Kaleta. Importantly, they could reproduce their findings in several independent cohorts of type-2 diabetic patients across Europe. Moreover, the bacterial species found to be major producers of agmatine were those known to be increased in the gut microbiome of metformin-treated type-2 diabetic patients.
Implications for metformin treatment
“Our results shed light on how the complex network of interactions between diet, microbiota and host impacts the efficacy of drugs,” says Cabreiro. “With our high-throughput screening approach we now finally have a tool at hand that allows us to tackle this complexity”. The findings of this study may help to inform dietary guidelines or the development of genetically engineered bacteria that could be used to enhance the beneficial effects of metformin. They may also provide a valuable insight into the evidence that suggests that metformin-treated type-2 diabetic patients are healthier and live longer than non-diabetic individuals.
Original publication:
Rosina Pryor, Povilas Norvaisas, Georgios Marinos , Lena Best, Louise Bruun Thingholm, Leonor Quintaneiro, Wouter De Haes, Daniela Esser, Silvio Waschina, Celia Lujan, Reuben L. Smith, Timothy Scott, Daniel Martinez-Martinez, Orla Woodward, Kevin Bryson, Matthias Laudes, Wolfgang Lieb, Riekelt H. Houtkooper, Andre Franke, Liesbet Temmerman, Ivana Bjedov, Helena M. Cochemé, Christoph Kaleta and Filipe Cabreiro (2019): Host-Microbe-Drug-Nutrient Screen Identifies Bacterial Effectors of Metformin Therapy. Cell Published August 29, 2019
DOI: 10.1016/j.cell.2019.08.003
Contact information:
Filipe Cabreiro
Host-Microbe Co-Metabolism Group
MRC London Institute of Medical Sciences (LMS)
Imperial College London
Contact: f.cabreiro@lms.mrc.ac.uk / +44 (0)20-8383-3768
Christoph Kaleta
Research Group Medical Systems Biology
Institute of Experimental Medicine
Kiel University / University Medical Center Schleswig-Holstein
Contact: c.kaleta@iem.uni-kiel.de / +49-(0)431-50030340
Weitere Informationen:
Host-Microbe Co-Metabolism Group
MRC London Institute of Medical Sciences (LMS), Imperial College London
Medical Systems Biology,
Institut für Experimentelle Medizin, Kiel University / UKSH
Cluster of Excellence „Precision Medicine in Chronic Inflammation“ (PMI),
Kiel University
Notes:
- Filipe Cabreiro’s work was supported by the Sir Henry Dale Fellowship from Wellcome and The Royal Society and the MRC-London Institute of Clinical Sciences at Imperial College London.
- Christoph Kaleta’s work was supported by the Collaborative Research Centre 1182 “Origin and Function of Metaorganisms” and the Cluster of Excellence “Precision Medicine in Chronic Inflammation” at Kiel University.
MRC London Institute of Medical Sciences is a core biomedical research Institute funded by the Medical Research Council, part of UK Research and Innovation. Based in West London at the Hammersmith Hospital, it is a vibrant research environment in which scientists, clinicians and patients collaborate to advance the understanding of biology and its application to medicine. The LMS pursues world-leading discovery science, and aims to address major health challenges (such as obesity, diabetes, heart disease and cancer), investigate the interplay between genes and environment, and understand the molecular basis of disease, development and ageing. LMS research programmes are supported in their delivery of world-class science by state-of-the-art facilities. LMS research is focused in three sections: Epigenetics, Genes and Metabolism, and Quantitative Biology.
The Collaborative Research Centre (CRC) 1182 “Origin and Function of Metaorganisms” is an interdisciplinary network of 80 researchers investigating associations between specific microbial communities and their multicellular hosts. Funded by the German Research Foundation (DFG), it is working on the issue of how plants and animals, including humans, form functional units (metaorganisms) together with highly specific microbial communities. The aim of the CRC 1182 is to understand why and how microbial communities enter into these long lasting connections with their hosts, and what functional consequences these interactions have. In total, the CRC 1182 includes researchers from five faculties at Kiel University, the GEOMAR Helmholtz Centre for Ocean Research Kiel, the Max-Planck-Institutes for Evolutionary Biology in Plön and for Marine Microbiology in Bremen.
The Cluster of Excellence “Precision Medicine in Chronic Inflammation” (PMI) is being funded from 2019 to 2025 through the German Excellence Strategy (ExStra). It succeeds the “Inflammation at Interfaces” Cluster, which was already funded in two periods of the Excellence Initiative (2007-2018). Around 300 members from eight institutions at four locations are involved: Kiel (Kiel University, University Medical Center Schleswig-Holstein (UKSH), Muthesius University of Fine Arts and Design, Kiel Institute for the World Economy (IfW), Leibniz Institute for Science and Mathematics Education (IPN)), Lübeck (University of Lübeck, University Medical Center Schleswig-Holstein (UKSH)), Plön (Max-Planck Institute for Evolutionary Biology) and Borstel (Research Center Borstel – Leibniz Lung Center). The goal is to translate interdisciplinary research findings on chronic inflammatory diseases of barrier organs to healthcare more intensively, as well as to fulfil previously unsatisfied needs of the patients. Three points are important in the context of successful treatment, and are therefore at the heart of PMI research: the early detection of chronic inflammatory diseases, the prediction of disease progression and complications, and the prediction of individual responses to treatment.