Building blocks for microbiome research
Kiel researchers, together with an international team, present a nematode model system as the basis for comprehensive microbiome research
All multicellular living beings are colonized by an unimaginably large number of microorganisms and have evolved together with these microbial symbionts from the beginning. The natural microbiome, i.e. the entirety of the bacteria, viruses and fungi that live in and on a body, is of central importance for the entire organism: it supports, for example, the absorption of nutrients, fends off pathogens and can also be involved in the development of serious diseases. Researchers use so-called model systems to unravel the highly complex interactions of host organisms and microorganisms, including for example, animal and plant species that can be easily studied in the laboratory, and a specific, simplified selection of colonizing microorganisms.
A research team from Kiel University, together with international experts from for example the University of California in Berkeley and the Baylor College of Medicine in Houston, has now developed a novel, improved model system for microbiome research, the so-called ‘Caenorhabditis elegans microbiome resource’ (CeMbio). It is based on a set of 12 bacteria that are representative of the natural composition of a nematode microbiome. This resource will allow more realistic microbiome studies in a variety of research fields, thereby taking advantage of the nematode C. elegans as an experimental system for microbiome research. The Kiel University research team from the Evolutionary Ecology and Genetics group headed by Professor Hinrich Schulenburg and the Medical Systems Biology group headed by Professor Christoph Kaleta recently presented the CeMbio resource and its main characteristics in the scientific journal G3: Genes Genomes Genetics.
A near-natural nematode microbiome
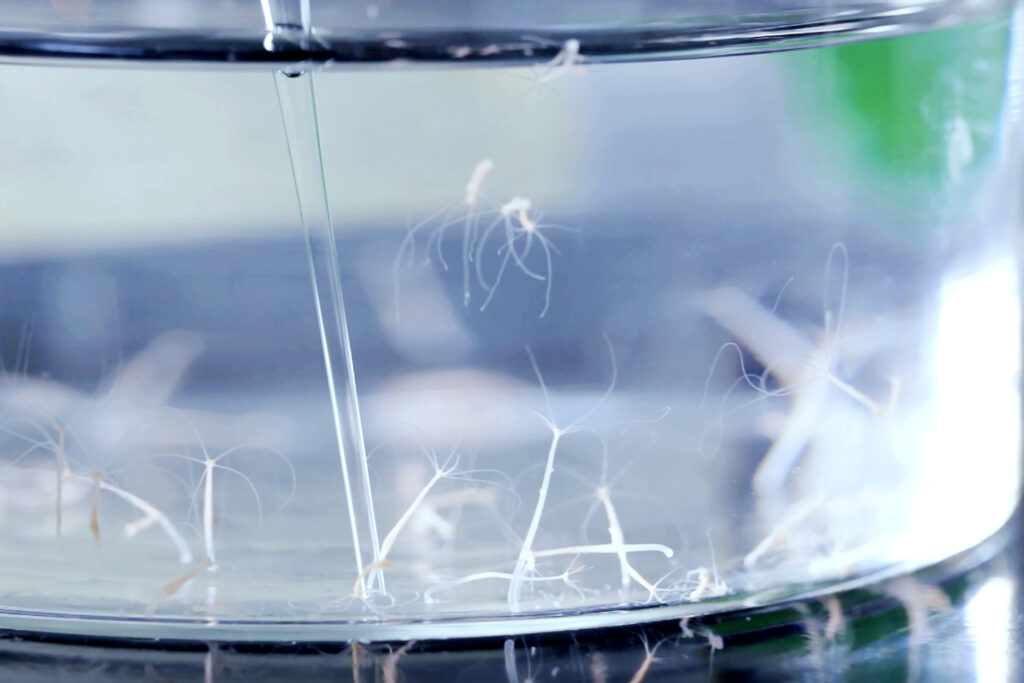
The starting point for the CeMbio project was previous work by the researchers from Kiel, in which they determined for the first time the naturally associated bacterial community of the nematode C. elegans, which is usually studied under sterile conditions. This enabled them to investigate the effects of the natural microbiome on the worm’s life functions and fitness. The components of the nematode microbiome that were identified in their entirety at that time were then narrowed down to twelve representative bacterial species. The scientists used them for colonization experiments of the nematode. “We transferred these bacteria individually in pure culture and in mixed cultures into germ-free worms and then observed their growth,” points out first author Dr Philipp Dirksen, former member of the Schulenburg group. “In this way, we were able to characterize the ability of individual bacteria to colonize the worm gut and the relationship of the different bacteria to each other in the natural host environment,” Dirksen continues.In a second part of the work, the scientists used whole genome sequences of all organisms present in the microbiome to reconstruct the metabolic networks and metabolic competences of the bacteria. Using bioinformatics methods, they were thus able to predict which metabolic products could be exchanged between host and microbiome. Overall, they thus created the basis for a realistic microbiome model for the detailed study of host-bacteria interactions in C. elegans.
An open source microbiome model
With the CeMbio model, the Kiel researchers are now offering a resource that is publicly accessible to colleagues in the C. elegans research community worldwide. In addition to the bacterial strains that can be easily cultivated under laboratory conditions, this includes their completely decoded genetic information as well as the complete modeling of the metabolic networks. The resource and generated database now provides the international research community with a novel toolkit that can be used to study numerous aspects of nematode biology from growth to development in the context of interactions with its natural microbiome. “Our CeMbio model opens the door for a more realistic characterization of a broad spectrum of interesting research questions ranging from disease development and the biology of aging to neurobiological aspects,” emphasizes the evolutionary biologist Schulenburg, head of the Kiel Evolution Center (KEC) at Kiel University. “The possibility of investigating the associated processes in the host’s interaction with its natural microbiome offers completely new perspectives for understanding the underlying molecular mechanisms,” continues Schulenburg.
In addition, the Kiel researchers hope that the new tool will also help to establish C. elegans more strongly than before as a model system for microbiome research. All in all, CeMbio provides scientists around the world with a novel resource to study this model nematode in its natural context. The work thus makes an important contribution to improving the overall understanding of the mechanisms by which the microbiome influences the health and disease of its host organism, as is being pursued by several research initiatives in Kiel, for example the Collaborative Research Center (CRC) 1182 “Origin and Function of Metaorganisms” or the Cluster of Excellence “Precision Medicine in Chronic Inflammation” (PMI).
Original publication:
Philipp Dirksen, Adrien Assié, Johannes Zimmermann, Fan Zhang, Adina-Malin Tietje, Sarah Arnaud Marsh, Marie-Anne Félix, Michael Shapira, Christoph Kaleta, Hinrich Schulenburg and Buck Samuel (2020): CeMbio – The Caenorhabditis elegans microbiome resource. G3: Genes Genomes Genetics DOI: 10.1534/g3.120.401309
Contact:
Prof. Hinrich Schulenburg
Evolutionary Ecology and Genetics research group,
Zoological Institute, Kiel University:
Phone: +49 (0) 431-880-4141
Email: hschulenburg@zoologie.uni-kiel.de
More information:
• Evolutionary Ecology and Genetics research group, Zoological Institute, Kiel University:
www.uni-kiel.de/zoologie/evoecogen
• „Kiel Evolution Center“ (KEC), Kiel University:
www.kec.uni-kiel.de
• Cluster of Excellence „Precision Medicine in Chronic Inflammation“ (PMI), Kiel University: www.precisionmedicine.de